What are scientific workflows?
Scientific workflows allow users to easily express multi-step computational tasks, for example retrieve data from an instrument or a database, reformat the data, and run an analysis. A scientific workflow describes the dependencies between the tasks and in most cases the workflow is described as a directed acyclic graph (DAG), where the nodes are tasks and the edges denote the task dependencies. A defining property for a scientific workflow is that it manages data flow. The tasks in a scientific workflow can be everything from short serial tasks to very large parallel tasks (MPI for example) surrounded by a large number of small, serial tasks used for pre- and post-processing.
| Reproducibility
Scientific workflows allow researchers to document and reproduce their analyses, ensuring their validity. |
Automation
Workflows automate repetitive and time-consuming tasks, thereby reducing the workload of researchers |
Scalability
Scale them to handle large data sets and complex analyses, enabling scientists for bigger research problems |
Reusability
Scale them to handle large data sets and complex analyses, enabling scientists for bigger research problems |
Pegasus overview
The Pegasus project encompasses a set of technologies that help workflow-based applications execute in a number of different environments including desktops, campus clusters, distributed computing environments such as PATh and OSG OSPool, supercomputing CI such as ACCESS and clouds. Pegasus bridges the scientific domain and the execution environment by automatically mapping high-level workflow descriptions onto distributed resources. It automatically locates the necessary input data and computational resources necessary for workflow execution. Pegasus enables scientists to construct workflows in abstract terms without worrying about the details of the underlying execution environment or the particulars of the low-level specifications required by the middleware (HTCondor, SLURM, or Amazon EC2). Pegasus also bridges the current cyberinfrastructures by effectively coordinating multiple distributed resources.
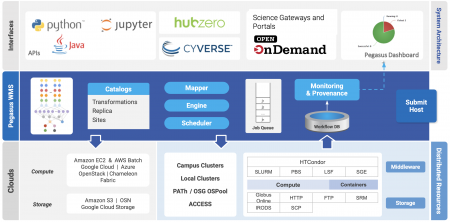
Pegasus has been used in a number of scientific domains including astronomy, bioinformatics, earthquake science , gravitational wave physics, ecology, electron microscopy(cryo-EM) and others. When errors occur, Pegasus tries to recover when possible by retrying tasks, by retrying the entire workflow, by providing workflow-level checkpointing, by re-mapping portions of the workflow, by trying alternative data sources for staging data, and, when all else fails, by providing a rescue workflow containing a description of only the work that remains to be done. It cleans up storage as the workflow is executed so that data-intensive workflows have enough space to execute on storage-constrained resources]. Pegasus keeps track of what has been done (provenance) including the locations of data used and produced, and which software was used with which parameters.
Pegasus has a number of features that contribute to its usability and effectiveness.
| Data Management
Pegasus handles data transfers, input data selection and output registration by adding them as auxiliary jobs to the workflow |
Error Recovery and Reliablity
Pegasus handles errors by retrying tasks, workflow-level checkpointing, re-mapping and alternative data sources for data staging. Debugging tools such as pegasus-analyzer help the user to debug the workflow in case of non-recoverable failures. |
Provenance Tracking
Pegasus allows users to trace the history of a workflow and its outputs, including information about data sources and softwares used |
Heterogeneous Environments
Pegasus can execute workflows in a variety of distributed computing environments such as HPC clusters, Amazon EC2, Google Cloud, Open Science Grid or ACCESS |
Others are listed below
- Portability / Reuse – User created workflows can easily be run in different environments without alteration. Pegasus currently runs workflows on top of HTCondor, distributed computing infrastructures such as PATh and OSG OSPool, supercomputing CI such as ACCESS, Amazon EC2, Google Cloud, and many campus clusters. The same workflow can run on a single system or across a heterogeneous set of resources.
- Performance – The Pegasus mapper can reorder, group, and prioritize tasks in order to increase overall workflow performance.
- Scalability – Pegasus can easily scale both the size of the workflow, and the resources that the workflow is distributed over. Pegasus runs workflows ranging from just a few computational tasks up to 1 million. The number of resources involved in executing a workflow can scale as needed without any impediments to performance.
- Containers Support – Pegasus has well defined support for application container technologies such as Docker and Singularity/Apptainer. This allows users to specify the container in which a particular job should run. Pegasus takes care of all the deployment and management of the container on the worker nodes as the workflow executes.
- Data Integrity – Pegasus does end to end automatic checksumming of workflow data to ensure data integrity. After every transfer checksums are matched to ensure data was not corrupted inadvertently during the data transfers.
Funding
Pegasus is funded by The National Science Foundation under Office of Advanced Cyberinfrastructure (OAC) grant #2138286. Previously, NSF has funded Pegasus under OAC SI2-SSI program grant #1664162, OCI SDCI program grant #0722019 and OCI SI2-SSI program grant #1148515.
Featured Publications
Deelman, E., Vahi, K., Rynge, M., Mayani, R., Ferreira da Silva, R., Papadimitriou, G., & Livny, M. (2019). The Evolution of the Pegasus Workflow Management Software. Computing in Science Engineering, 21(4), 22–36. https://doi.org/10.1109/MCSE.2019.2919690
Deelman, E., Vahi, K., Juve, G., Rynge, M., Callaghan, S., Maechling, P. J., Mayani, R., Chen, W., Ferreira da Silva, R., Livny, M., & Wenger, K. (2015). Pegasus: a Workflow Management System for Science Automation. Future Generation Computer Systems, 46, 17–35. https://doi.org/10.1016/j.future.2014.10.008
59,998 views